好的颜值,人人都爱,是你接触有趣的灵魂的敲门砖。单细胞数据分析也是如此,人人都知道需要降维聚类分群。
有了好的代码,甚至非本专业的财务人员都可以复制粘贴我们写好的的代码,参考前面的例子:人人都能学会的单细胞聚类分群注释 , 但不一定每个人都能合理的解释各个单细胞亚群,而标记基因是其中最重要的一个手段来辅助说明你的细胞亚群。广为人知的seurat包就提供了5个方法来进行标记基因可视化,让我们来总结整理一下吧。
先介绍seurat最常用的示例数据
基本上每个人开始学习单细胞,都是从这个文档开始:https://satijalab.org/seurat/articles/pbmc3k_tutorial.html
假如你安装GitHub的包没有问题,就直接使用下面的代码:
library(Seurat)
# devtools::install_github('satijalab/seurat-data')
library(SeuratData)
library(ggplot2)
library(patchwork)
library(dplyr)
InstallData("pbmc3k")
data("pbmc3k")
pbmc3k
pbmc=pbmc3k
如果GitHub无法访问,也可以
# Load the PBMC dataset
# https://cf.10xgenomics.com/samples/cell/pbmc3k/pbmc3k_filtered_gene_bc_matrices.tar.gz
pbmc.data <- Read10X(data.dir = "filtered_gene_bc_matrices/hg19/")
# Initialize the Seurat object with the raw (non-normalized data).
pbmc <- CreateSeuratObject(counts = pbmc.data, project = "pbmc3k", min.cells = 3, min.features = 200)
pbmc
这个 pbmc 变量就是全部单细胞数据分析所需要的,是一个被精心定义好的对象哦:
> pbmc
An object of class Seurat
13714 features across 2700 samples within 1 assay
Active assay: RNA (13714 features, 0 variable features)
走单细胞基础流程即可
所以单细胞转录组的数据分析的核心,就在于前面的 An object of class Seurat 制作:
有了做好的Seurat 对象,后续代码非常标准,如下:
pbmc[["percent.mt"]] <- PercentageFeatureSet(pbmc, pattern = "^MT-")
#Visualize QC metrics as a violin plot
VlnPlot(pbmc, features = c("nFeature_RNA", "nCount_RNA", "percent.mt"), ncol = 3)
# FeatureScatter is typically used to visualize feature-feature relationships, but can be used for anything calculated by the object, i.e. columns in object metadata, PC scores etc.
plot1 <- FeatureScatter(pbmc, feature1 = "nCount_RNA", feature2 = "percent.mt")
plot2 <- FeatureScatter(pbmc, feature1 = "nCount_RNA", feature2 = "nFeature_RNA")
plot1 + plot2
pbmc <- subset(pbmc, subset = nFeature_RNA > 200 & nFeature_RNA < 2500 & percent.mt < 5)
pbmc <- NormalizeData(pbmc, normalization.method = "LogNormalize", scale.factor = 1e4)
pbmc <- FindVariableFeatures(pbmc, selection.method = 'vst', nfeatures = 2000)
# Identify the 10 most highly variable genes
top10 <- head(VariableFeatures(pbmc), 10)
# plot variable features with and without labels
plot1 <- VariableFeaturePlot(pbmc)
plot2 <- LabelPoints(plot = plot1, points = top10, repel = TRUE)
plot1 + plot2
pbmc <- ScaleData(pbmc, vars.to.regress = "percent.mt")
pbmc <- RunPCA(pbmc, features = VariableFeatures(object = pbmc))
#
pbmc <- FindNeighbors(pbmc, dims = 1:10)
pbmc <- FindClusters(pbmc, resolution = 0.5)
# Look at cluster IDs of the first 5 cells
head(Idents(pbmc), 5)
table(pbmc$seurat_clusters)
pbmc <- RunUMAP(pbmc, dims = 1:10)
# note that you can set `label = TRUE` or use the LabelClusters function to help label individual clusters
DimPlot(pbmc, reduction = 'umap')
VlnPlot(pbmc, features = c("MS4A1", "CD79A"))
FeaturePlot(pbmc, features = c("MS4A1", "GNLY", "CD3E", "CD14", "FCER1A", "FCGR3A", "LYZ", "PPBP", "CD8A"))
new.cluster.ids <- c("Naive CD4 T", "CD14+ Mono", "Memory CD4 T", "B", "CD8 T", "FCGR3A+ Mono", "NK", "DC", "Platelet")
names(new.cluster.ids) <- levels(pbmc)
pbmc <- RenameIdents(pbmc, new.cluster.ids)
DimPlot(pbmc, reduction = 'umap', label = TRUE, pt.size = 0.5) + NoLegend()
走完基础流程后这个pbmc对象就很丰富的信息啦:
> pbmc
An object of class Seurat
13714 features across 2638 samples within 1 assay
Active assay: RNA (13714 features, 2000 variable features)
2 dimensional reductions calculated: pca, umap
我们会得到如下所示的分群情况:
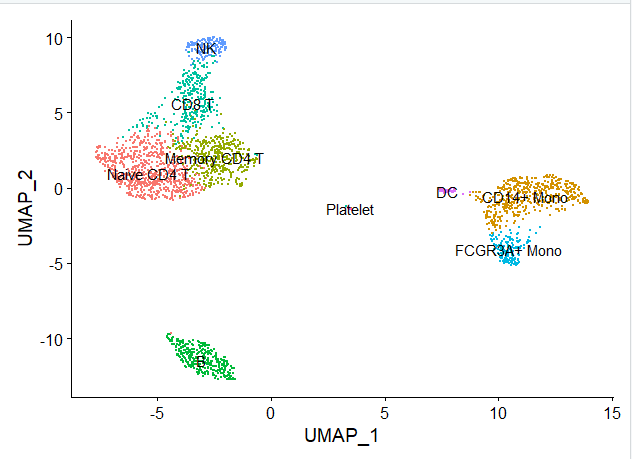
根据生物学背景知识,我们需要可视化如下所示的各个单细胞亚群的标记基因,如下所示: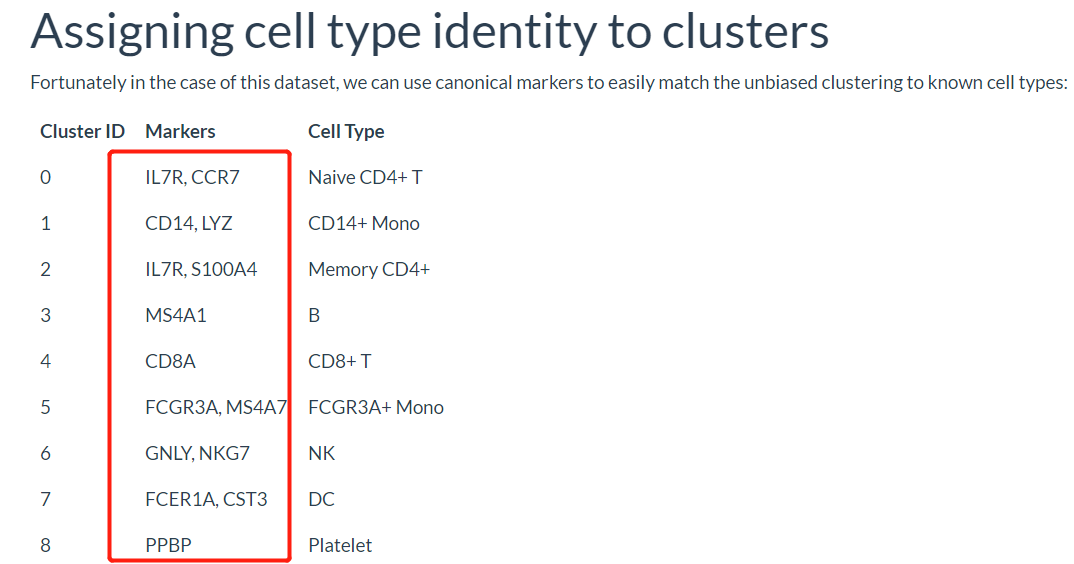
这个时候有5个可视化方法,分别是:小提琴图,坐标映射图,峰峦图,气泡图,热图。全部的代码如下所示:
# https://satijalab.org/seurat/archive/v3.0/visualization_vignette.html
VlnPlot(pbmc, features = c("MS4A1", "CD79A"))
FeaturePlot(pbmc, features = c("MS4A1", "CD79A"))
RidgePlot(pbmc, features = c("MS4A1", "CD79A"), ncol = 1)
features= c('IL7R', 'CCR7','CD14', 'LYZ', 'IL7R', 'S100A4',"MS4A1", "CD8A",
'FCGR3A', 'MS4A7', 'GNLY', 'NKG7','FCER1A', 'CST3','PPBP')
DotPlot(pbmc, features = unique(features)) + RotatedAxis()
DoHeatmap(subset(pbmc, downsample = 100), features = features, size = 3)
都是超级简单的,封装好了的,只需要具备基础R语言知识,就可以做出像模像样的美图哦!
小提琴图
如下所示:
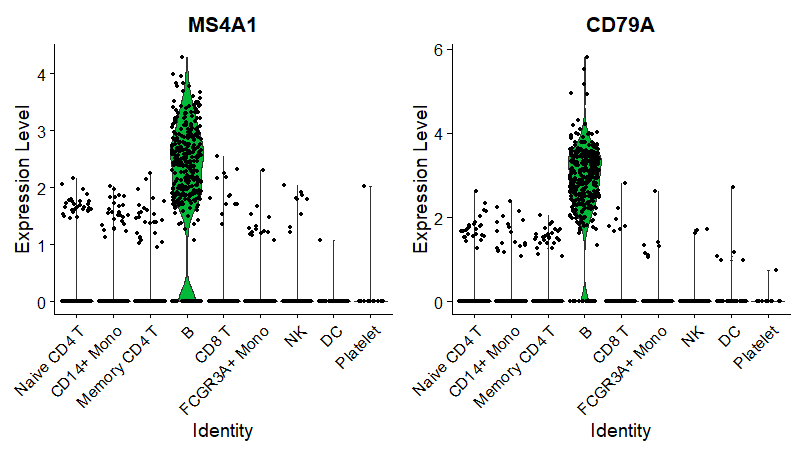
坐标映射图
如下所示:
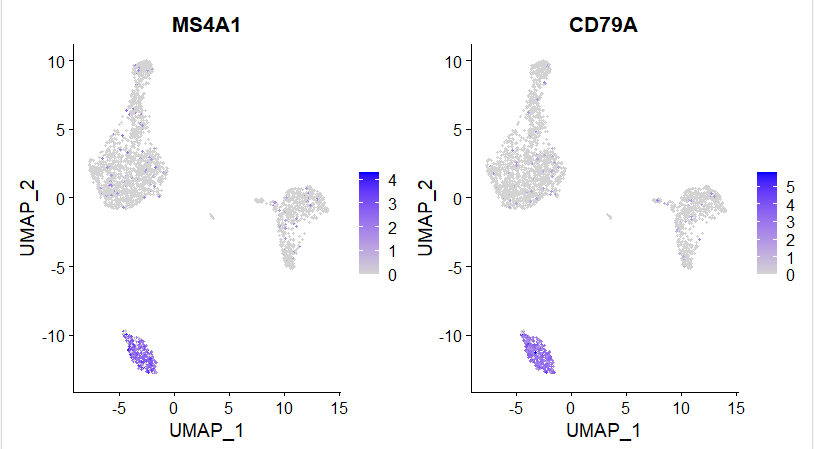
峰峦图
如下所示:
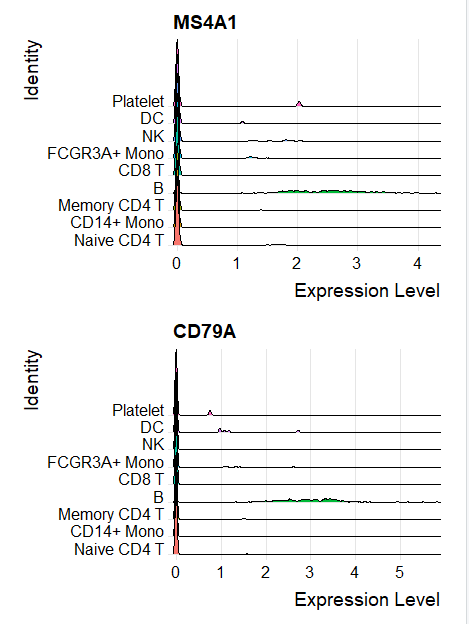
气泡图
如下所示:
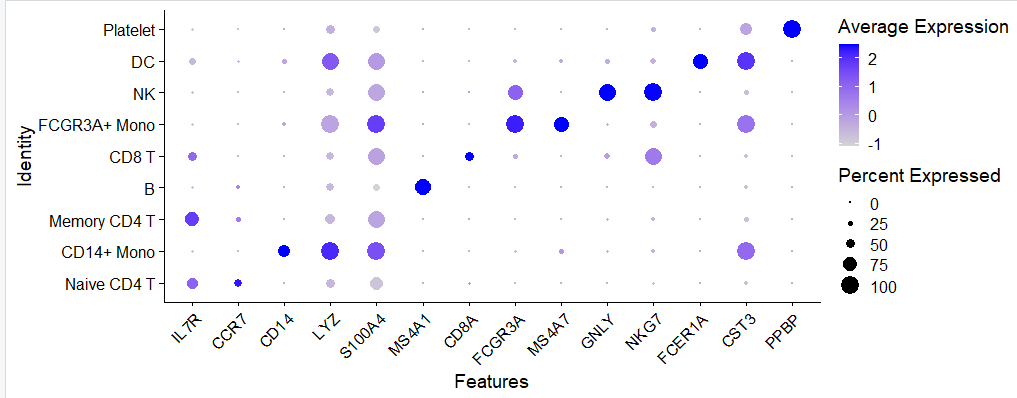
热图
如下所示:
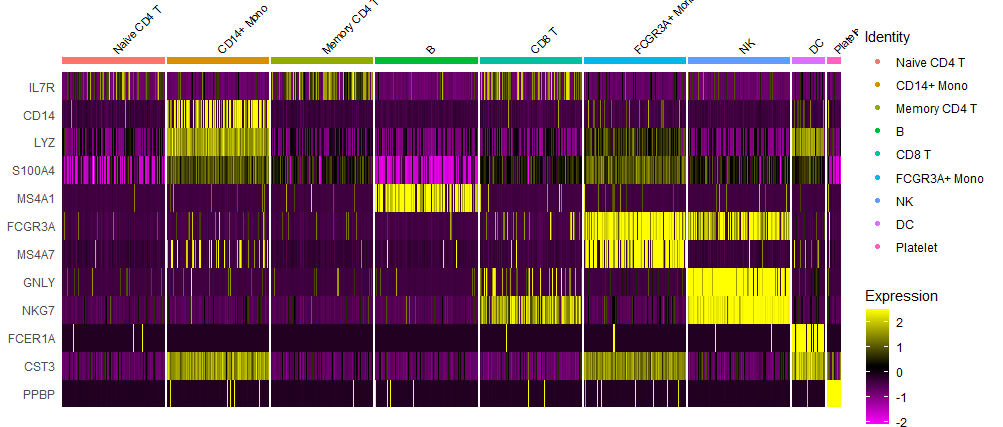
文末小调研
这5个可视化方法,小提琴图,坐标映射图,峰峦图,气泡图,热图。你最喜欢哪个?